LIGSITE
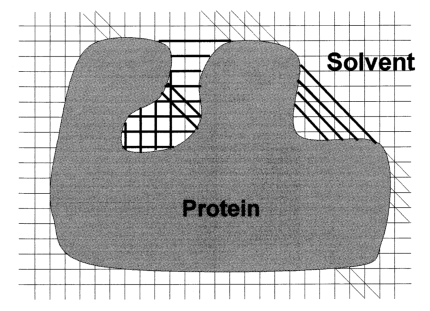
In LIGSITE, pockets are identified by scanning along the x, y, and z axes and the cubic diagonals (only four diagonals are shown) for areas that are enclosed on both sides by protein (indicated by boldface lines):
In LIGSITE, pockets are identified by scanning along the x, y, and z axes and the cubic diagonals (only four diagonals are shown) for areas that are enclosed on both sides by protein (indicated by boldface lines):
Essentially, it identifies solvent-protein-solvent events to predict pockets and cavities that may be binding sites. The accuracy with which the surface of pockets and cavities is determined depends on the step size that is used. At an infinitely small step size, the surface determined by this algorithm would be equivalent to the contact surface. A large grid step size, e.g. 2.0 Å, decreases calculation time but results in a very angular surface.
PASS
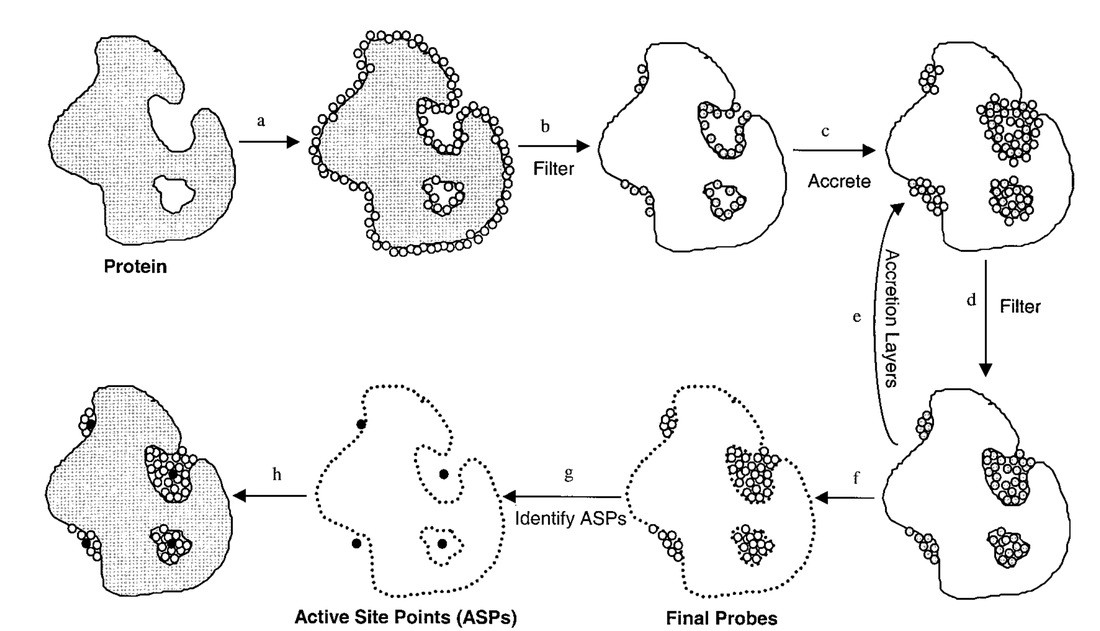
PASS (Putative Active Sites with Spheres) uses the concept of probe spheres and “burial scores” to identify binding sites:
Note: Steps 1-8 correspond to a-h
PASS (Putative Active Sites with Spheres) uses the concept of probe spheres and “burial scores” to identify binding sites:
Note: Steps 1-8 correspond to a-h
- Coat the protein with an initial layer of spherical probes
- These probes are filtered to eliminate those that:
- Clash with the protein
- Are not sufficiently buried
- Lie within 1 Å of a more buried probe.
- A new layer of spheres is accreted on top of the previous layer.
- These probes are filtered as described in step 2.
- A new layer of spheres is accreted onto the existing probes, as in step 3.
- Accretion and filtering (steps 5 and 4) are repeated until a layer occurs in which no newly found probes survive the filters. This leaves the final set of probe spheres.
- Probe weights (PW) are computed for each sphere and active site points (ASPs) are then identified from amongst the final probes.
- The final PASS prediction is produced. The final probe spheres are first smoothed, leaving only clusters of four or more.
SURFNET
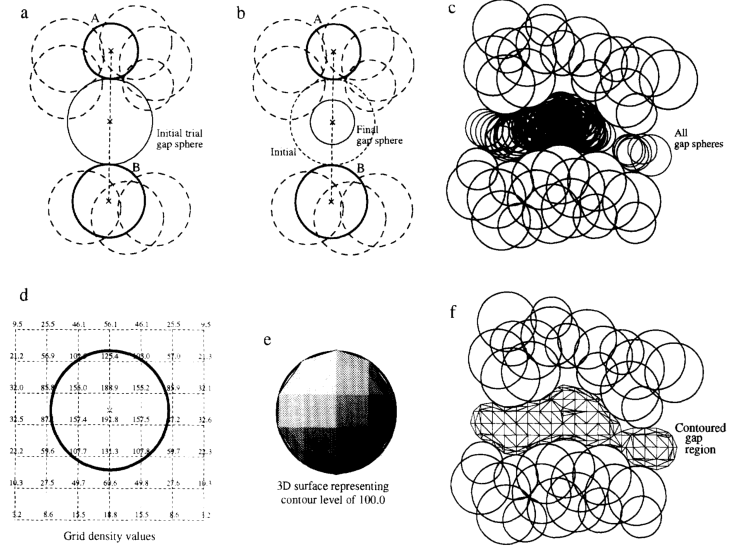
SURFNET uses “gap spheres” to fill regions between two molecules in order to identify pockets/cavities. The gap regions are filled with gap spheres that are defined by the surface of two specific atoms. Then, the gap spheres are reduced in size until they no longer intersect the volumes of other atoms in the area. This process is repeated for all combinations of two atoms within the gap region. Then, a polyhedron approximating the sphere can be used to generate a larger polyhedron representing the gap region as a whole:
SURFNET uses “gap spheres” to fill regions between two molecules in order to identify pockets/cavities. The gap regions are filled with gap spheres that are defined by the surface of two specific atoms. Then, the gap spheres are reduced in size until they no longer intersect the volumes of other atoms in the area. This process is repeated for all combinations of two atoms within the gap region. Then, a polyhedron approximating the sphere can be used to generate a larger polyhedron representing the gap region as a whole:
Roll
The key concept of Roll is to generate a crust-like surface called the probe surface enveloping protein. This allows Roll to identify the region between the probe surface and protein surface as a ‘pocket’ and the region surrounded by protein surface as a ‘cavity’. Such a probe surface is created by “rolling” a probe sphere around the surface of the protein. The size of the probe sphere can determine the size of the binding sites to be identified (a smaller probe sphere may cause some binding sites not to be identified).
The key concept of Roll is to generate a crust-like surface called the probe surface enveloping protein. This allows Roll to identify the region between the probe surface and protein surface as a ‘pocket’ and the region surrounded by protein surface as a ‘cavity’. Such a probe surface is created by “rolling” a probe sphere around the surface of the protein. The size of the probe sphere can determine the size of the binding sites to be identified (a smaller probe sphere may cause some binding sites not to be identified).
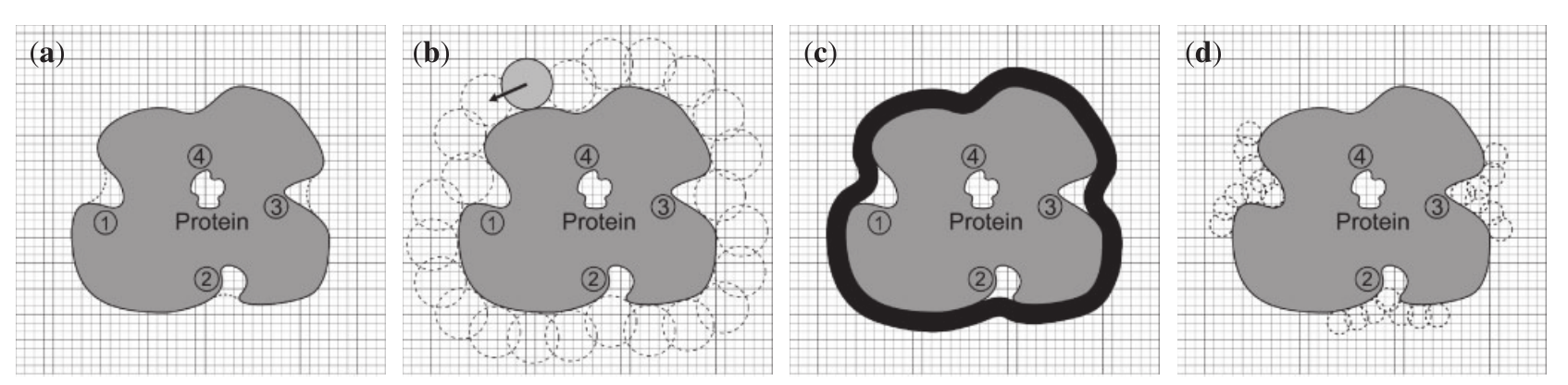
Note: 1-4 corresponds to a-d
- 2D Slice of the grid system. The dashed and solid lines show the probe surface and the protein surface, respectively. The grey region is the protein. Regions 1–3 are defined as pockets and region 4 is defined as a cavity.
- The rolling process. The light grey ball indicates the starting position and the dashed balls show the trace of rolling.
- The black area is the probe surface. Pockets 1-3 have been identified (between probe surface and protein surface) in addition to cavity 4 (surrounded by protein surface).
- The smaller probe sphere causes pocket 1 to disappear and pockets 2–3 to become smaller, while it does not affect cavity 4.